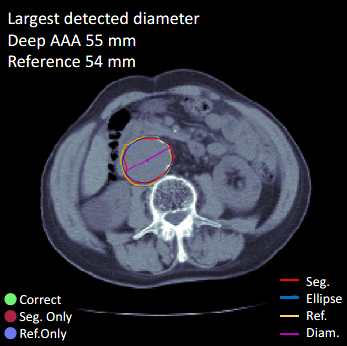
An abdominal aortic aneurysm is a potentially serious condition that can be missed by radiologists when it was not the primary reason for a scan. We discuss in detail how Nuance and CCDS are collaborating to use deep learning approaches for detecting them.
About the author: Rupert Brooks is a Senior Research Scientist in Nuance Global Research and Development focused on problems relating to diagnostic imaging that affect the longer term development of Nuance’s AI marketplace. His current research interests include deep learning models for medical imaging, federated learning and privacy preserving approaches to machine learning. He has previously done research in real time surgical simulation and image guidance for radiotherapy. Rupert obtained a Ph.D. in Electrical and Computer Engineering from McGill University, currently holds an affiliate assistant professor position at Concordia University and is a member of the Order of Engineers of Quebec.
Deep learning based approaches have recently made great strides in automating some types of medical image analysis. Despite some hype about deep learning algorithms replacing radiologists, it is most likely that deep learning algorithms will become sophisticated tools for radiologists, assisting and complementing their expertise. AI based approaches may assist by improving workflow efficiency, providing confirmation, or remaining alert for unusual circumstances which may escape the radiologist’s attention.
Introduction
The aorta is the largest artery in the body. It connects to the left ventricle, loops over the heart, and then descends roughly vertically to the pelvis, where it splits into the two iliac arteries. Freshly oxygenated blood from the lungs is pumped through it with each heartbeat. Aneurysms, swellings which indicate a weakening of the vessel walls of the aorta, occur with moderate frequency as people age. So long as the aneurysm remains intact, symptoms, if any, are usually minor. However, rupture of an abdominal aortic aneurysm (AAA) can lead to massive internal bleeding and is frequently fatal.
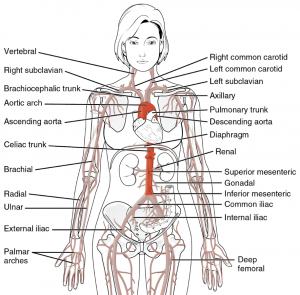
There are a range of treatment options if the aneurysm is noticed before it ruptures. However, as the symptoms are minimal, it is frequently detected only incidentally, when an examination is being done for some other reason. Thus when a radiologist reads a scan covering the aorta, they should report the presence of any aneurysm. However, the literature shows that these are often missed. Together with the researchers at the Massachusetts General Hospital (MGH) Center for Clinical Data Science (CCDS), we see an opportunity for an AI based approach to provide a “second set of eyes” to assist radiologists in incidentally detecting this condition.
CT Scans
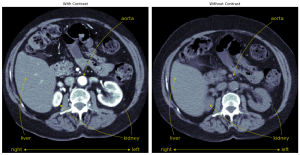
The most likely scan type which would permit incidentally finding an abdominal aortic aneurysm is a CT (computed tomography) scan. A CT scanner works by rotating an X-ray source around the patient, projecting a beam of X-rays through them at different angles. The set of projections can be mathematically combined to form a 3D image of the body. To enhance the contrast of the blood vessels, a contrast agent may be injected into the bloodstream. The blood vessels will absorb more x-rays, and appear brighter, when the contrast is present. While the presence of contrast will definitely make an abdominal aortic aneurysm easier to detect, to be effective our approach should work with or without contrast.
The patient is usually scanned lying down on a motorized platform, which moves them through the rotating ring containing the x-ray source. The resulting CT scan is usually structured as a series of slices perpendicular to the head to foot axis of the patient. These slices may be spaced closer together or further apart depending on the scan parameters. To be practical, a method for detecting abdominal aortic aneurysms must work with a variety of slice spacings. While there are a number of additional variables that can change the imaging process, contrast agent and slice spacing are the two most important variables the method must contend with.
Approach
In the first phase of this project, we have concentrated on the abdominal region of the aorta. In this area, a healthy aorta has a fairly straight cylindrical form. To detect aneurysms we first must segment the aorta from the scan, and then measure it to detect any bulges which would indicate an aneurysm. We chose to apply a deep learning approach to the segmentation problem, and a more classical geometric approach to the problem of determining if the aorta is healthy.
Segmentation of the aorta
For the segmentation problem, we chose to use the U-Net neural network architecture, a state of the art approach to segmentation of medical imagery. The U-Net structure can be described at a high level as an encoder/decoder network with skip connections. Specifically, both the encoder and decoder sections are constructed of a series of convolutional blocks. Each block in the encoder section increases the number of channels in the feature map, and decreases spatial detail. Conversely, each block in the decoder section reduces the number of channels and increases the spatial detail. The output of each encoder block before reducing the resolution is branched off and combined with the decoder part after increasing its resolution. The structure is usually drawn with the encoder descending on the left and the decoder rising on the right, in a U shape, which gives rise to its name. Conceptually, semantic information (the what) is carried through the bottleneck in the filters, while spatial information (the where) is carried across to the decoder network through the skip connections.
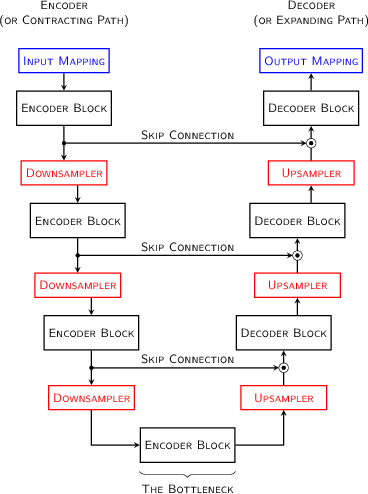
Originally developed in a 2D context, several variations of the U-Net have been explored which extend the idea into 3D. In our case, we expect a wide variation in slice spacing along the head-foot axis, but anticipate reasonably consistent and high resolution in the other directions. In this case, to allow operation across variable numbers of slices, we did not reduce the spatial resolution in the head/foot direction. This allows the system to process scans with an arbitrary number of slices.
We obtained 321 CT scans from 223 unique patients at the Massachusetts General Hospital to use for developing the network. The aorta was manually segmented by CT technologists to provide a reference segmentation. The dataset was split roughly 80/10/10 into training validation and test sets. The network was trained by directly optimizing the Dice score between the output segmentation and the reference segmentation. The weights which performed best on the validation set were chosen. For further details on the design or training procedure, consult this paper.
Identification of aneurysms
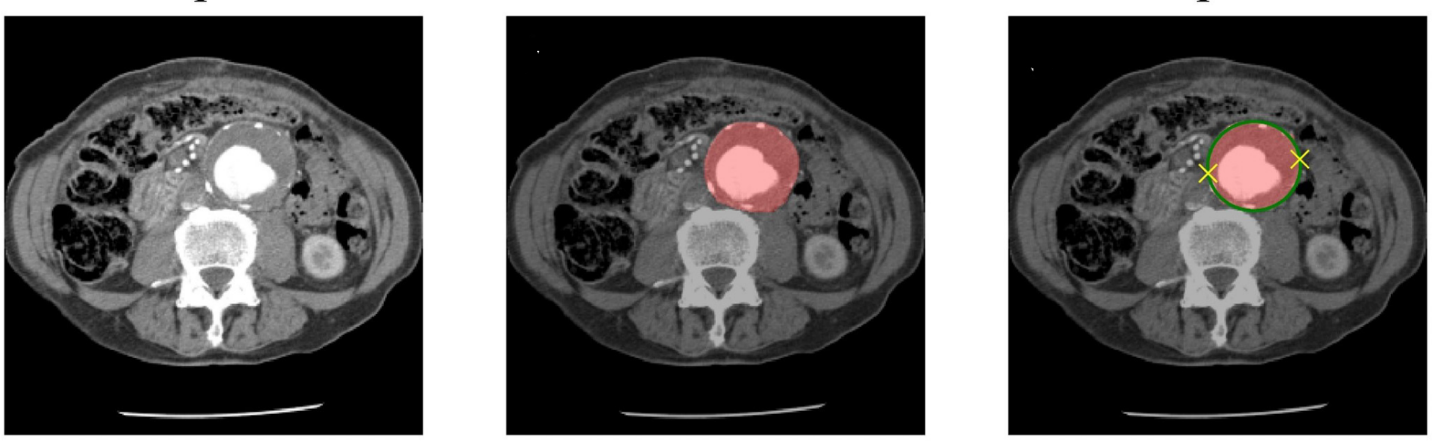
Once trained, the network proposes a segmentation of the aorta for any input CT image of the abdomen. This segmentation is then measured to determine if it shows a bulge which would be interpreted as an aneurysm. A classical geometric approach was used for this, an ellipse is fitted to each axial slice. We consider the major axis of this ellipse to be a good estimator of the maximum diameter of the aorta in this slice, and we correct the estimate for the tilt of the centerline. Clinical guidelines suggest that a diameter greater than 3.0 cm should be considered as an aneurysm.
Results
On the original dataset of 321 scans, the output of the model could be compared to the reference segmentations. The estimated maximum diameter could also be compared to the maximum diameter of the reference segmentation produced by the technologists. However, while the official diagnosis was presumably consistent with the reference segmentation size, it was not directly tracked. In a slightly unexpected, but very promising result, the system performed as well on scans with contrast as on those without. The segmentation algorithm achieved an average Dice score of 0.90, and had an average error with respect to the maximum diameter of the reference segmentation of only -2 mm.
As a further test, an additional 57 scans were obtained – these scans did not have reference segmentations, but had clinically validated diagnoses of the presence or absence of an abdominal aortic aneurysm, and estimates of the aneurysm size made by medical experts in a clinical context. On this additional, more clinically relevant test, the system also performed well, with a mean delta of less than a millimeter, a sensitivity of 85% and a specificity of 100%.
The system has difficulty however, in correctly segmenting the aorta in the thorax. This is not surprising, as we concentrated on the abdomen for this development and did not have thorax data in the training data. For the purpose of this study, thorax data was trimmed out of the second dataset by hand and the results presented here correspond to that trimmed dataset. For realistic clinical use, however, a means of handling the thoracic aorta needs to be developed. Many scans intended for the abdomen contain a certain amount of the thorax, and, more importantly, aneurysms of the thoracic aorta also occur, and it is important to detect them as well.
Conclusion and Future work
By combining a U-net based segmentation with a geometric analysis, our system is able to detect abdominal aortic aneurysms in the abdomen with high sensitivity and specificity. These results compare favorably with the performance of human radiologists in detecting incidental findings as described here. While these numbers aren’t directly comparable, not least because they are on different datasets, it shows the approach has potential to be useful.
This work was intentionally limited to the abdominal region. Work to extend the technique to the thorax is currently in progress. In the thorax the shape of the aorta is more complex, it loops up and over the heart and is surrounded by other blood vessels. This presents additional challenges both to the segmentation approach, and to the geometric post processing used to identify aneurysms.
The work so far has also been limited to a research context on retrospective data. Nuance and the CCDS are currently working to integrate the system with the Nuance AI Marketplace, and eventually to make it available in a clinical context. Achieving clinical utility and regulatory acceptance is challenging, but necessary in order for the work to deliver its potential benefits.
Acknowledgements
This system described in this post was developed at the Massachusetts General Hospital Center for Clinical Data Science, and includes the work of multiple contributors. Jen-Tang Lu was the lead author of this work, with contributions from Rupert Brooks, Stefan Hahn, Jin Chen, Varun Buch, Gopal Kotecha, Katherine P. Andriole, Brian Ghoshhajra, Joel Pinto, Paul Vozila, Mark Michalski, and Neil A. Tenenholtz.
All usage of patient data in this project was approved by the institutional review board of the Massachusetts General Hospital.
Publication and press
This work has been published at the [22nd International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI 2019). A preprint is available on Arxiv.
An early version of the system was presented at the 2019 Society for Imaging Informatics in Medicine (SIIM) conference, where it won the Best Scientific Paper award.
An early demonstration of the system integrated with the Nuance AI marketplace was conducted at RSNA 2018.